
- support
- info@evidentic.com
- +49 (0) 30 959 99 8831
Host cell proteins (HCPs) are unwanted proteins produced by the host organism during the manufacturing and production of biologics. HCPs, also vary apropos to the type and strain of the host organism. Although HCPs are generally found in small quantities (parts per million expressed as nanograms per milligrams of the desired recombinant protein), they are not approved to be a part of the final biologics. But, the removal of HCPs is time-consuming and expensive. HCPs are present at different stages of the manufacturing and differ in their composition and abundance depending on four factors such as host expression system, the manner in which the biologic of interest is expressed, the purification process, and the biologic molecule being expressed.
Since HCPs are produced from non-human expression systems, they have the potential to induce an immune response in humans. Even though the magnitude and nature of the immune response vary with respect to the composition, a number of foreign components introduced, and the individual immune response. Furthermore, such risks can be difficult to analyze and evaluate because of the limited information from pre-clinical and clinical studies. Moreover, HCP’s may reduce the efficacy of the biotherapeutic of interest through their proteolytic activity. Hence, characterization and purification of HCP’s are of prime importance.
To meet regulatory criteria, biologics developers must verify that HCPs are in the acceptable range and consider factors such as:
Generally acceptable, and most biologic products contain 10-1000 ppm HCPs, with most current processes resulting in HCP levels less than 100 ppm. Whereas, in the case of monoclonal antibodies (mAbs) production, typically achieving less than 100 ppm total HCP has been considered ideal.

Hence, HCP levels in the drug is considered a critical quality attribute (CQA). Therefore, unerring detection, identification and quantification of HCPs during the biologics development is critical in ensuring patient safety, and is a regulatory requirement. To be amenable with such stipulations, the application of advanced purification and analytical methods is an absolute necessity.
The first step towards the removal of HCPs in your drug is the characterization or profiling of HCPs based on the selected expression system (for example, E. coli or CHO cell lines) and downstream processes. Once the qualitative analysis is performed, the HCPs are quantified at various steps during product development and purification, in order to track and trace the amounts of HCP in the final product (Figure). The two main technology platforms used to evaluate the drug HCP profile are discussed here.
ELISAs are colorimetric tests used to quantify biomolecular interactions. This immunoassay is used to detect HCPs in biologics as they are easy to implement, fast, and economically viable providing good precision and sensitivity, making the assay compliant in a QC setting and for commercial use. Commonly, the test is conducted in a 96 or 384 well microplate. Among the 4 different available formats of ELISA, a sandwich ELISA is generally preferred over others to detect HCPs, which are usually present in the parts per million levels. During this procedure, low levels of HCPs are added to bind to the antibody-coated on microplate. Subsequently, HCPs are bound with a reporting antibody tagged with an enzyme or fluorescence to amplify the HCPs. In this format, the amount of binding antibodies that can be present in the microplate is usually <1 mg/well. Hence, depending on antibody diversity, the amount of each anti HCP can be determined.
Despite the advantages, the commercial ELISA test is not sensitive to artifacts present in complex biologics, and does not provide an accurate estimate of total HCPs for specific processes. Therefore, HCP-specific ELISAs are developed in-house for better characterization and analysis of the HCP profile of the product (Figure). But, production of these customized HCP-specific ELISAs also requires considerable development effort and long lead times to allow for production and characterization of custom immunoreagents derived from animal sources, along with validation of satisfactory HCP coverage. Moreover, such crucial customized immunoreagents have a limited shelf-life and must be reproduced and re-validated periodically to support ongoing analysis of the marketed product. Another important factor that demands ELISA methods to utilize a host-cell specific assay to detect HCPs from the appropriately matched cell source is the significant differences in the proteomes of distinct host types. Also, host cell protein expression is affected by culture conditions and methods, as well as other aspects of living cells, such as age. Practically, the generic ELISA only provides about 75% coverage of HCPs, hence, is only effective as a primary assay for initial development around 50% of the time. Another common problem present in ELISA-based assays is that many production hosts produce impurities that are closely related to the desired biological product. Moreover, orthogonal approaches, such as 2D gel, Western blot and mass spectrometry (MS) analysis, is recommended in combination with ELISAs in order to identify individual HCP species. Furthermore, to reduce the laborious and time-consuming nature of ELISA, commercial quantitation ELISA assays with automation protocols have been developed.
Recently, the usage of advanced biosensors has put a high-throughput twist on ELISAs and has shown to increase HCP’s detection range, speed and sensitivity. Such immunosensors generate a measurable signal provided by a transducer. These immunosensors are categorized based on their transduction mode as:
(1) Optical
(2) Electrochemical and
(3) Piezoelectric devices
Mass spectrometry (MS) analysis is an orthogonal or supporting method to ELISA for HCP analysis, rather than a QC and product release method. Although MS-based technologies require a higher initial cost and a high level of technical expertise, once developed and optimized for a particular expression system, it can be used routinely to aid appropriate process development (Figure). With high accuracy and sensitivity, MS can specifically identify different HCP profiles and distinguish process-derived impurities, as well as quantify the components to a single-digit ppm value. Usually, a phased approach is used for HCP analysis with regard to MS. First, the HCPs are identified, followed by quantification assays.
Phase I involves qualitative detection of impurities from samples taken at various steps in the downstream process. Hence, in the context of HCP discovery and monitoring, liquid chromatography coupled with mass spectrometry (LC-MS) is used to achieve the goal. A sample of the biologic culture (e.g., cell lysate or supernatant) from each phase of the downstream process step is first denatured with specific chemicals or enzymes. The sample is then subjected to LC separation and MS analysis. There are two approaches to MS analysis:
DIA is gaining popularity as it enables the quantification of 1000s of proteins with low variation and high reproducibility. Nevertheless, a combination of the two approaches is often employed to ensure a broad-range characterization of HCPs.
However, the analytical workflow can vary depending on the type of LC and MS methods utilized. Reverse-phase LC (RPLC) is the most preferred sample separation method. Lately, in order to improve sensitivity, a two-dimensional LC (2D-LC) approach has garnered attention, wherein the sample is passed through two columns. 2D-LC utilizes a combination of various LC platforms, such as cation exchange chromatography (SCX)- RPLC, strong anion exchange chromatography (SAX)-RPLC, size exclusion chromatography (SEC)-RPLC, hydrophilic interaction chromatography (HILIC)-RPLC, RPLC-capillary zone electrophoresis (CZE), and RPLC-RPLC. The method used can be either a comprehensive 2D-LC, where the entire sample is passed through the first and the second column; or it can be heart-cutting 2D-LC, where the sample is separated using the first column and selected subsets are passed through the second column to separate co-eluting compounds. Although 2D-LC presents a higher resolving power, on the downside, it considerably increases the run-time and demands more sample amounts.
Another strategy that has been developed to improve sensitivity is the addition of molecular weight cut-off enrichment prior to LC-MS. In this case, the biologic drug is filtered out from the solution, leaving behind low abundance proteins below the MW cut-off value. This makes it easier to characterize low MW HCPs and also reduces the dynamic range across proteins in the sample by 1000-fold. Applying this method to reference drugs, recent studies have reported the presence of a number of HCPs that were previously not described. They successfully demonstrated the detection of low MW HCPs (11 – 17 kDa) that were present at very low concentrations (0.1 – 8 ppm) in the sample.
In Phase II, relative quantification is performed against a set of standard proteins (Figure). Here, a set of supplementary proteins at known concentrations are added to the sample to achieve a quantification value relative to those proteins. In the final Phase III, quantification is done against heavy-labelled marker peptides. In this heavy-isotope labelled analogues of HCPs that are identified in phases I and II, are added at known concentrations to the sample. This will help to improve accuracy by comparing values of native versus the marker peptides.
HCPs being an important CQA, it is imperative to identify and remove all the HCPs that can cause immunogenicity and affect the potency and stability of the recombinant biologic drug or the excipients in the formulation. In this regard, host-cell line engineering is an important and upcoming strategy. Using multiplex-CRISP/Cas9 gene knock-out technology, scientists have developed a “clean CHO” cell line that can produce fewer HCPs. This approach can be used to delete genes coding for proteins that can affect product quality, and hence will help ease the downstream process. Furthermore, null cells can be used as HCP reference standards to identify antigens and to develop process-specific assays. The null cell can be the parental cell in the case of a mammalian cell line, or a cell transfected with an empty vector.
To sum up, product samples must be analyzed throughout product development and purification steps, to identify and monitor the presence of HCPs. Once identified, HCPs can be quantified by using a relative quantification method, by evaluating against protein standards added to the sample. And more importantly, a simple workflow is always preferred keeping in mind the aspect of reproducibility.
Due to the intrinsic heterogeneity of the host cell proteome, it is challenging to identify and remove all the unwanted host-cell proteins. As per the regulatory agencies, the maximum permitted level of total HCP in biologic drugs is 100 ppm or 100 nanograms per mg. Nevertheless, certain HCPs can degrade product quality even at very low concentrations.
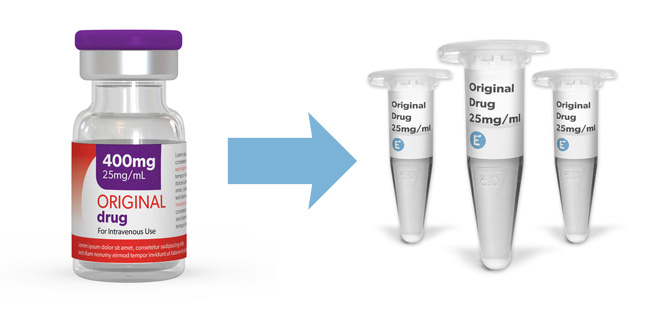
Do you require clinical-grade molecules for your early drug development programs? At Evidentic we source commercially EU-approved biologics as research consumables. We do not produce the molecules, instead, repackage the original vial content into small and convenient aliquots. Just browse our portfolio and order the quantity that you need.
Discover clinical-grade molecules


Evidentic GmbH
Martin-Buber-Str. 10
14163 Berlin